Installation
Requirements
QuPath (Steps 1–3)
- QuPath 0.6.0+
- Cellpose extension installed in QuPath — configure the Cellpose and Cellpose SAM Python paths in
Edit → Preferences → Cellpose/Omnipose
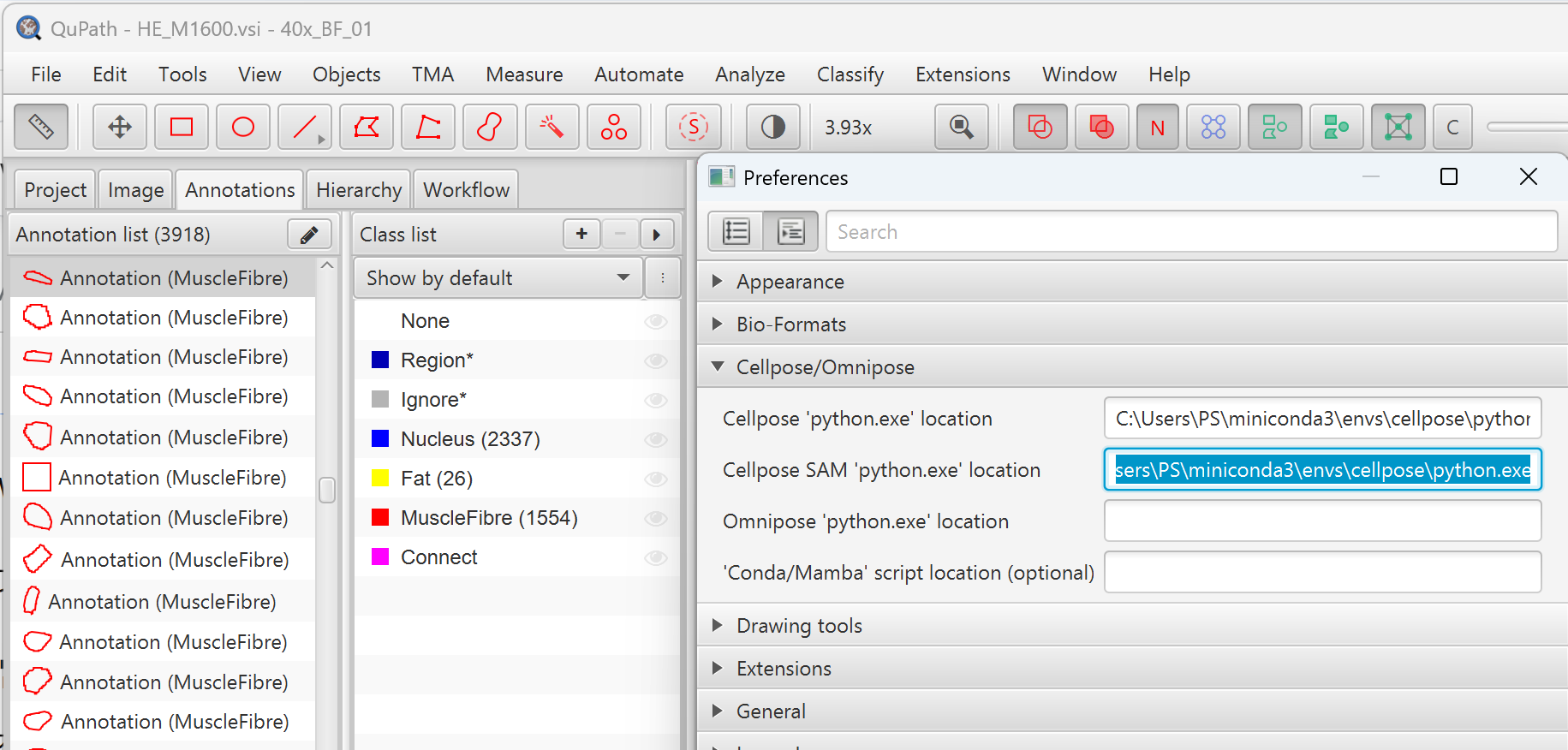
- GPU recommended for Cellpose segmentation
- A pre-trained pixel classifier named
"Fat in muscle", along with all Groovy scripts and Python scripts, can be downloaded from this GitHub repository
Python (Step 4)
- Python 3.9+
- Conda environment recommended
Download
1. Clone or download
MyoPath/
├── MyoPath1_roi.groovy
├── MyoPath2_segment.groovy
├── MyoPath3_export.groovy
├── MyoPath4_analysis.py
├── project.qpproj # QuPath project file
├── classifiers/
│ ├── classes.json
│ └── pixel_classifiers/
│ ├── Fat in muscle.json # Required pixel classifier
│ └── ... # Other optional classifiers
├── data/ # QuPath slide entries
├── exports/ # Step 3 output (per-sample)
├── results/ # Step 4 aggregated output
└── src/
├── data_loader.py
├── coordinate_processor.py
├── tissue_analyzer.py
├── visualizer.py
└── dystrophy_analyzer.pyPixel Classifiers
The classifiers/pixel_classifiers/ directory is part of the QuPath project and contains pre-trained pixel classifiers in JSON format. The pipeline requires a classifier named Fat in muscle for fat infiltration detection in Step 2. If you are setting up a new QuPath project, you need to train this classifier yourself (see Requirements) or copy it from an existing project.
2. Set up Python environment
conda create -n myopath python=3.10 -y
conda activate myopath
pip install tiatoolbox numpy scipy matplotlib shapely pandas3. Verify installation
python -c "import tiatoolbox, numpy, scipy, matplotlib, shapely, pandas; print('All dependencies OK')"Troubleshooting
Cellpose TileFile null error in Step 2
The downsample factor is too small, causing oversized tiles. The default downsample: 10.0 should work. If you still see this error, increase the value (e.g., 12.0 or 15.0).
No muscle fibers detected
- Ensure the ROI covers actual muscle tissue
- Check that Cellpose and its Python environment are properly installed
- Try lowering
cellprobThreshold(e.g., from 0 to −2)
Python ModuleNotFoundError
Make sure you run from the MyoPath/ directory, or use --input to specify the exports path.
Conda An unexpected error has occurred after Step 4
An unexpected error has occurred. Conda has prepared the above report.This is a known conda 25.7.0 bug on Windows with GBK encoding. It does not affect result generation — your output files are produced correctly despite the error message. You can safely ignore it, or suppress it with:
CONDA_NO_PLUGINS=true conda run -n myopath python MyoPath4_analysis.py --allOut of memory during batch processing
Reduce --cores to limit parallel processes (each sample uses ~2–4 GB RAM).