MyoPath
Under Development
MyoPath is currently under active development and not yet publicly available.
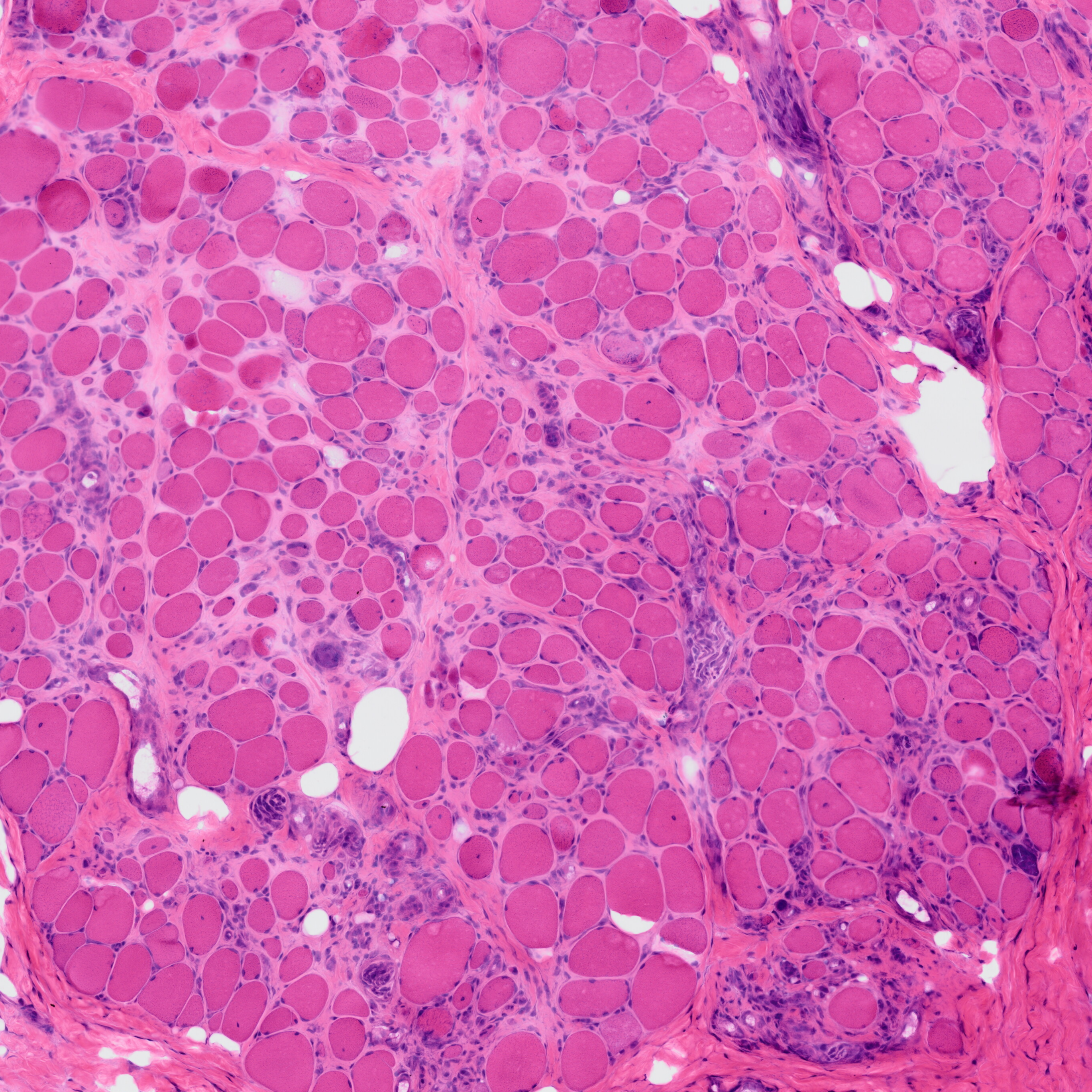
MyoPath is a deep learning pipeline for objective morphometric assessment of skeletal muscle biopsies from routine H&E-stained whole slide images (WSI).
ROI Region
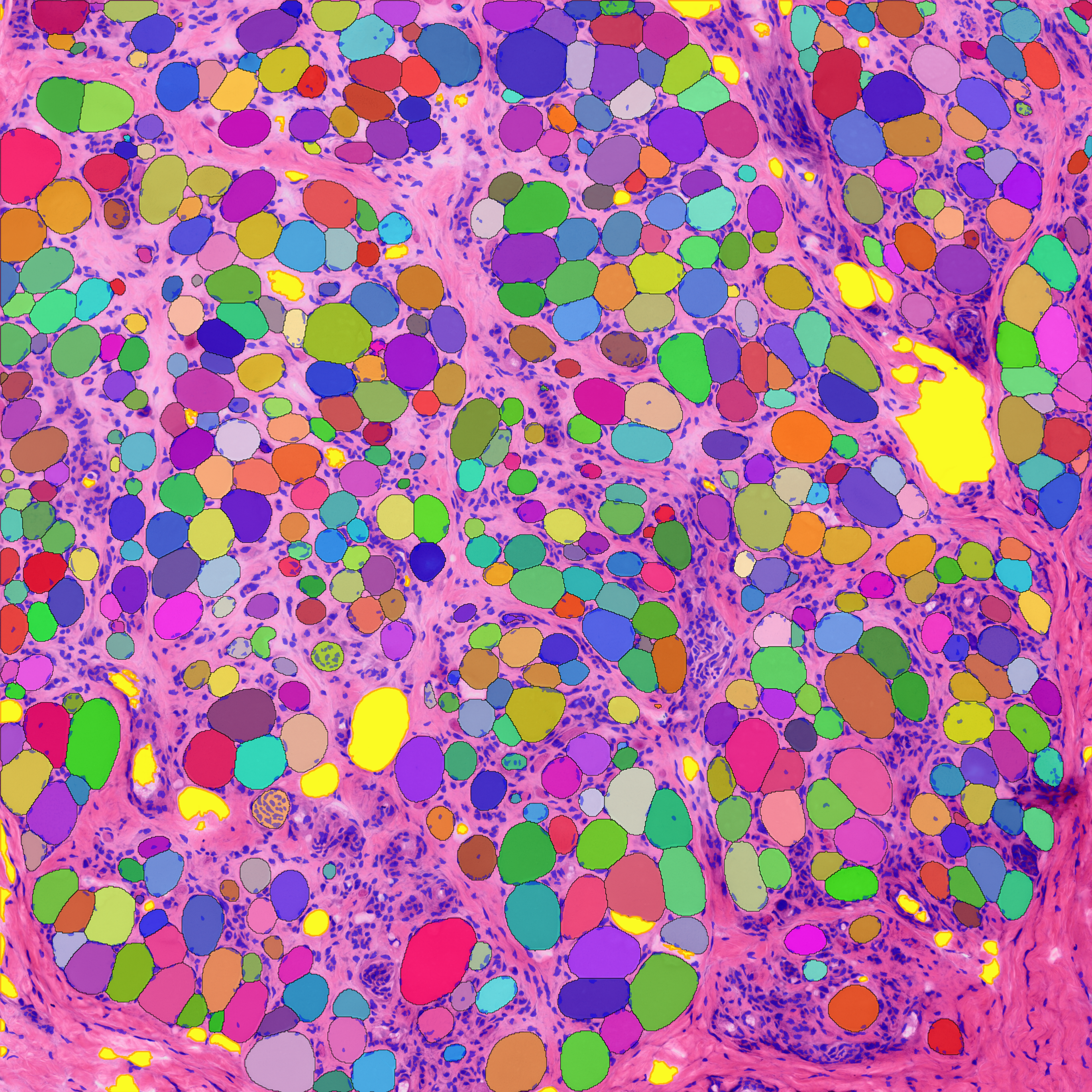
Muscle Fibers
Fat Regions
Connective Tissue
Fiber Size Distribution
Fiber Shape
Nuclei Analysis
Pathology Indicators
Image Comparison
Overview
MyoPath implements a four-tissue segmentation pipeline — Cellpose-SAM for myofiber instance segmentation, a pixel classifier for fat infiltration, watershed detection for nuclei, and Boolean subtraction for connective tissue — to extract 37 morphometric features per sample. From these, seven clinically interpretable pathology indicators are derived, with Nuclear Centralization Index (NCI) and Fiber Size Variability Coefficient (Fiber CV) serving as primary biomarkers.
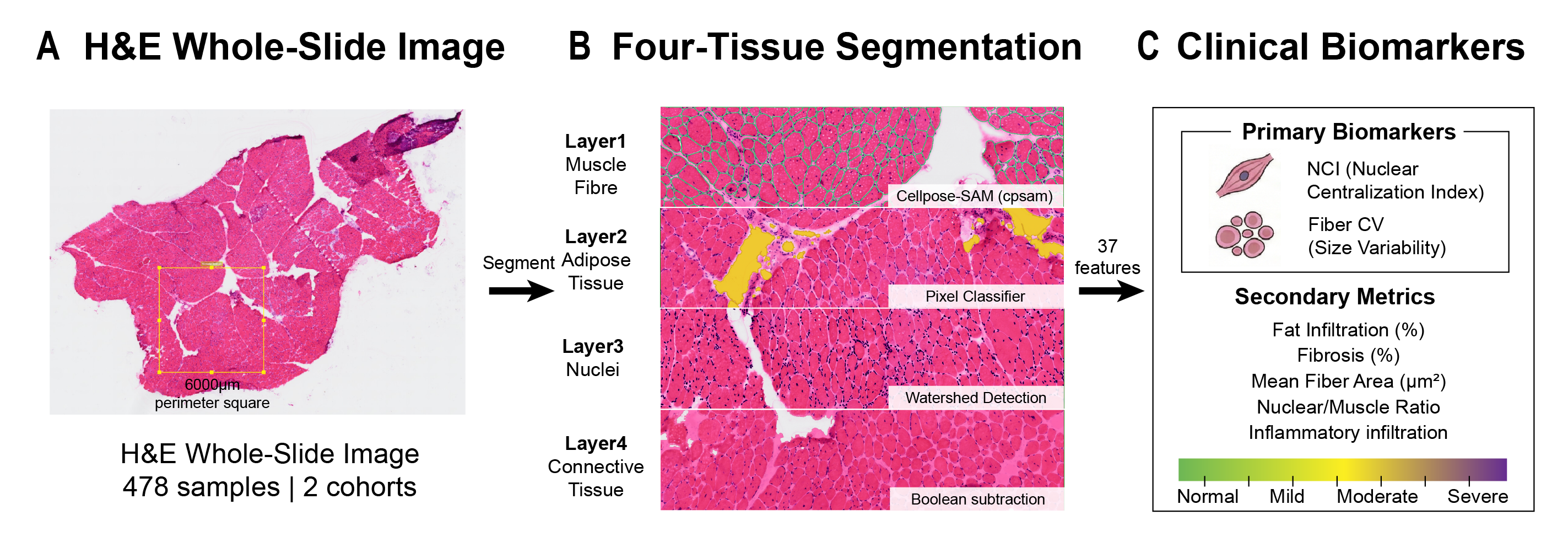
The pipeline has been validated on 478 H&E whole-slide images from two independent cohorts (HuashanMuscle, n = 79; GTEx, n = 399).
Key Features
- Four-tissue segmentation: Myofiber, fat, nucleus, and connective tissue from a single H&E section
- 37 morphometric features across five biological categories (details)
- Seven pathology indicators covering nuclear positioning, fiber size, shape, tissue composition, and cellular reaction
- MyoPath Score: Composite severity measure (AUC = 0.873 on external validation)
- Primary biomarkers: NCI (p = 1.3 x 10⁻⁵) and Fiber CV (p = 2.9 x 10⁻⁴)
Segmentation Pipeline
| Layer | Tissue | Method | Dice |
|---|---|---|---|
| 1 | Myofiber instances | Cellpose-SAM | 0.92 |
| 2 | Fat infiltration | Pixel classifier | 0.95 |
| 3 | Nuclei | Watershed detection | 0.87 |
| 4 | Connective tissue | Boolean subtraction | 0.88 |
Citation
Zhong H*, Gao M*, et al. MyoPath: A Deep Learning Pipeline for Objective Morphometric Assessment of Skeletal Muscle Biopsies. Manuscript in preparation.